Cross-Modality Domain Adaptation for Medical Image Segmentation
Unsupervised 3D Semantic Segmentation Domain Adaptation
👋 Join crossMoDA 2023! The crossMoDA 2021 paper accepted in Medical Image Analysis!

Aim
Domain Adaptation (DA) has recently raised strong interests in the medical imaging community. By encouraging algorithms to be robust to unseen situations or different input data domains, Domain Adaptation improves the applicability of machine learning approaches to various clinical settings. While a large variety of DA techniques has been proposed, most of these techniques have been validated either on private datasets or on small publicly available datasets. Moreover, these datasets mostly address single-class problems. To tackle these limitations, the crossMoDA challenge introduced the first large and multi-class dataset for unsupervised cross-modality Domain Adaptation. Compared to the previous crossMoDA instance, which made use of multi-institutional data acquired in controlled conditions for radiosurgery planning and focused on a 2 class segmentation task (tumour and cochlea), the 2023 edition extends the segmentation task by including multi-institutional, heterogenous data acquired for routine surveillance purposes and introduces a sub-segmentation for the tumour (intra- and extra-meatal components) thereby leading to a 3 class problem.
Task
Task: Intra- and extra-meatal vestibular schwannoma and cochlea segmentation
The goal of the segmentation task is to segment two key brain structures (tumour and cochlea) involved in the follow-up and treatment planning of vestibular schwannoma (VS). The segmentation of these tumour and cochlea structures is required for radiosurgery, a common VS treatment, and is also key to improve routine surveillance. Anatomically, the tumour region can be divided into two sub regions, called intra- and extra-meatal regions, corresponding to being inside or outside the inner ear canal respectively. The guidelines for reporting results in vestibular schwannoma mention that the intra-meatal and extra-meatal portions of the tumour must be distinguished, and the largest extra-meatal diameter should be used to report the size of the tumour. Moreover, the size and volume features extracted from the extra-meatal region are considered as the most sensitive radiomic features for the evaluation of VS growth. The diagnosis and surveillance in patients with VS are commonly performed using contrast-enhanced T1 (ceT1) MR imaging. However, there is growing interest in using non-contrast imaging sequences such as high-resolution T2 (hrT2) imaging, as it mitigates the risks associated with gadolinium-containing contrast agents.
For this reason, we proposed an unsupervised cross-modality segmentation benchmark (from ceT1 to hrT2) that aims to perform the intra- and extra-meatal regions of VS and cochlea segmentation on hrT2 scans automatically. The training source and target sets are respectively unpaired annotated ceT1 and non-annotated hrT2 scans from both pre-operative and post-operative time points. To validate the robustness of the proposed approaches on different T2 settings, multi-institutional, heterogeneous scans from centres in the UK and Tilburg, NL are used in this task.
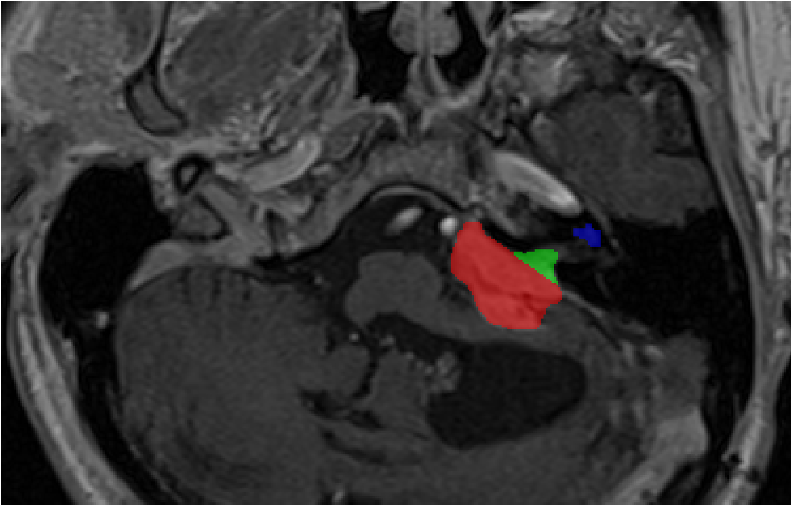 Source (contrast-enhanced T1)
Source (contrast-enhanced T1)
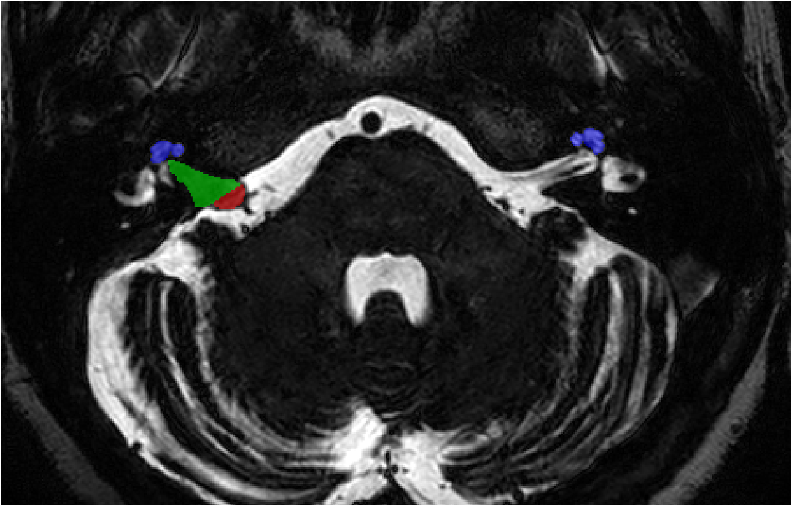 Target (high resolution T2)
Target (high resolution T2)
Data
London SC-GK data
All images were obtained on a 32-channel Siemens Avanto 1.5T scanner using a Siemens single-channel head coil:
- Contrast-enhanced T1-weighted imaging was performed with an MPRAGE sequence with in-plane resolution of 0.4×0.4mm, in-plane matrix of 512×512, and slice thickness of 1.0 to 1.5 mm (TR=1900 ms, TE=2.97 ms, TI=1100 ms)
- High-resolution T2-weighted imaging was performed with a 3D CISS or FIESTA sequence in-plane resolution of 0.5x0.5mm, in-plane matrix of 384x384 or 448x448, and slice thickness of 1.0 to 1.5 mm (TR=9.4 ms, TE=4.23ms).
Tilburg SC-GK data
All images were obtained on a Philips Ingenia 1.5T scanner using a Philips quadrature head coil:
- Contrast enhanced T1-weighted imaging was performed with a 3D-FFE sequence with in-plane resolution of 0.8×0.8mm, in-plane matrix of 256×256, and slice thickness of 1.5 mm (TR=25 ms, TE=1.82 ms).
- High-resolution T2-weighted imaging was performed with a 3D-TSE sequence with in-plane resolution of 0.4x0.4mm, in-plane matrix of 512×512, and slice thickness of 1.0 mm (TR=2700 ms, TE=160 ms, ETL=50).
All data will be made available online with a permissive non-commercial copyright-license (CC BY-NC-SA 4.0), allowing for data to be shared, distributed and improved upon. All structures were manually segmented in consensus by the treating neurosurgeon and physicist using both the ceT1 and hrT2 images. To cite this data, please refer to https://arxiv.org/abs/2201.02831.
UK MC-RC data:
Images were obtained on various scanners from SIEMENS, Philips, General Electrics, Hitachi MRI scanners with the magnetic field strengths 1.0T/1.5T/3.0T. Slice thickness, voxel volume and intensities vary significantly across all ceT1 weighted and T2 weighted imaging (see here for more details).
Rules
No additional data is allowed, including the data released on TCIA and pre-trained models. The use of a generic brain atlas is tolerated as long as its use is made clear and justified.
Example of tolerated use cases:- Spatial normalisation to MNI space
- Use of classical single-atlas based tools (e.g., SPM)
Examples of cases that are not allowed:
- Multi-atlas registration based approaches in the target domain
No additional annotations are allowed.
Models can be adapted (trained) on the target domain (using the provided target training set) in an unsupervised way, i.e. without labels.
The participant teams will be required to release their training and testing code and explain how they fine-tuned their hyper-parameters. Note that the code can be shared with the organisers only as a way to verify validity, and if needed, NDAs can be signed.
The top 3 ranked teams will be required to submit their training and testing codes in a docker container for verification after the challenge submission deadline in order to ensure that the challenge rules have been respected.
Evaluation
Classical semantic segmentation metrics, in this case, the Dice Score (DSC) and the Average Symmetric Surface Distance (ASSD), will be used to assess different aspects of the performance of the region of interest. These metrics are implemented here. The metrics (DSC, ASSD) were chosen because of their simplicity, their popularity, their rank stability, and their ability to assess the accuracy of the predictions.
Participating teams are ranked for each target testing subject, for each evaluated region (i.e., iIntra-meatal VS, extra-meatal VS and cochlea), and for each measure (i.e., DSC and ASSD). The final ranking score for each team is then calculated by firstly averaging across all these individual rankings for each patient (i.e., Cumulative Rank), and then averaging these cumulative ranks across all patients for each participating team.
Timeline
| Release of the training and validation data. | |
| Start of the validation period. | |
| 1st July 2023: | Start of the evaluation period. |
| 10th July 2023: | End of the evaluation period. |
| 8th October 2023: | Challenge results are announced at MICCAI 2023. |
| 30th November 2023: | Participants are invited to submit their methods to the MICCAI 2023 BrainLes Workshop. |
| December 2023: | Submission of a joint manuscript summarizing the results of the challenge to a high-impact journal in the field. |
Winners of crossMoDA-2022 challenge
Methods of the top-performing 2022 teams are described here.
Task 1: Segmentation
| # | Team Name | Affiliation | DSC | Technical report |
|---|---|---|---|---|
| 1 | ne2e | Peking University, Beijing, China | 86.9 | Unsupervised Domain Adaptation in Semantic Segmentation Based on Pixel Alignment and Self-Training (PAST) |
| 2 | MAI | Korea University, Korea | 86.8 | Multi-view Cross-Modality MR Image Translation for Vestibular Schwannoma and Cochlea Segmentation |
| 3 | LaTIM | Inserm, France | 85.9 | Tumor blending augmentation using one-shot generative learning for vestibular schwannoma and cochlea cross-modal segmentation |
Task 2: Koos Classification
| # | Team Name | Affiliation | MA-MAE | Technical report |
|---|---|---|---|---|
| 1 | SJTU_EIEE_2 | Shanghai Jiao Tong University, China | 0.26 | Koos Classification of Vestibular Schwannoma via Image Translation-Based Unsupervised Cross-Modality Domain Adaptation |
| 2 | Super Polymerization | Radboud University, the Netherlands | 0.37 | Unsupervised Cross-Modality Domain Adaptation for Vestibular Schwannoma Segmentation and Koos Grade Prediction based on Semi-Supervised Contrastive Learning |
| 3 | skjp | Muroran Institute of Technology, Japan | 0.84 | Unsupervised Domain Adaptation for MRI Volume Segmentation and Classification Using Image-to-Image Translation |